The nucleus is a large,
membranous organelle found only in eukaryotes. Its presence
makes possible eukaryote-only processes, such as
post-transcriptional RNA processing (capping,
polyadenylation, editing, splicing).
(Splicing
is no longer a eukaryote-only process!)
|
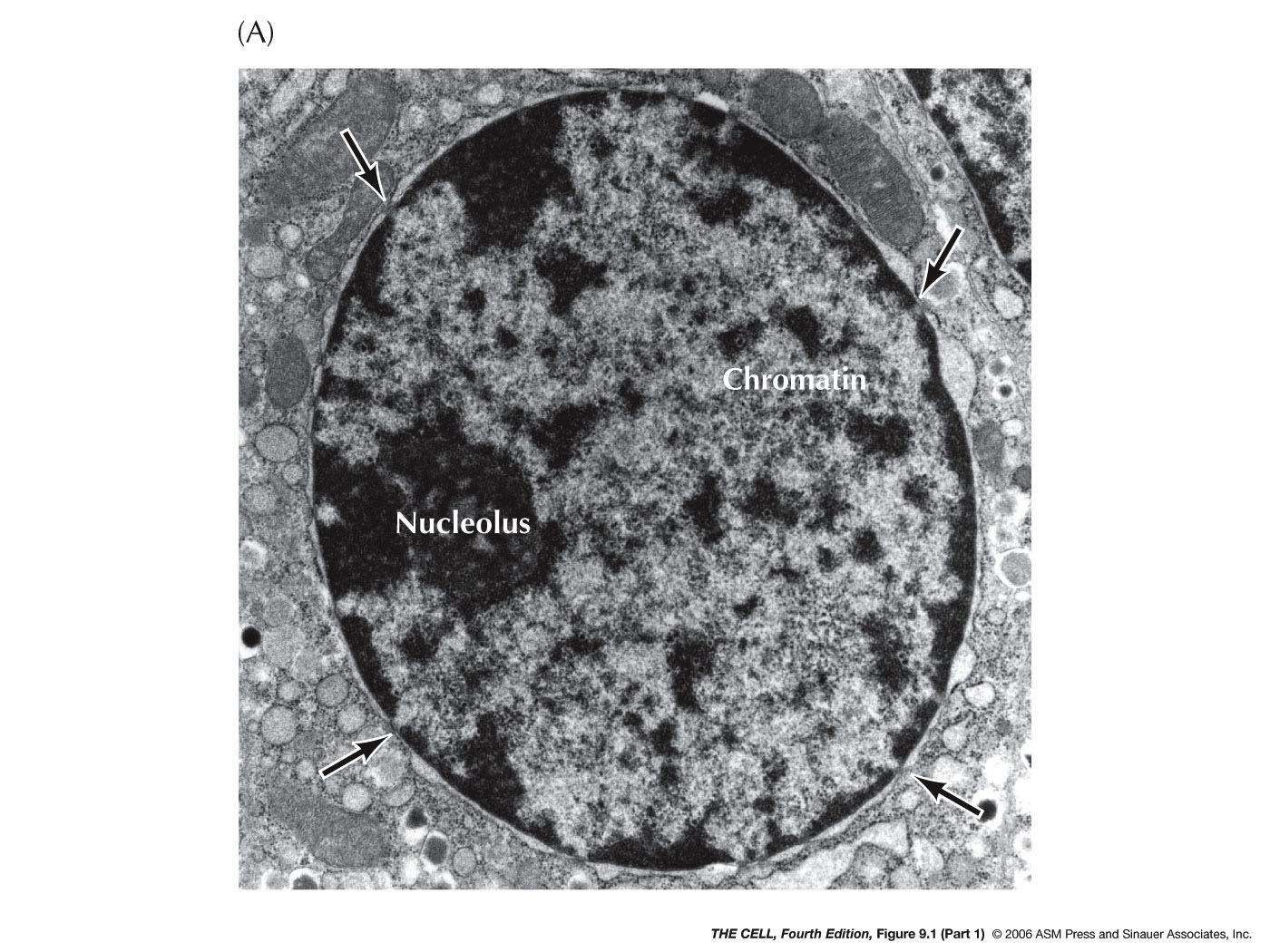 |
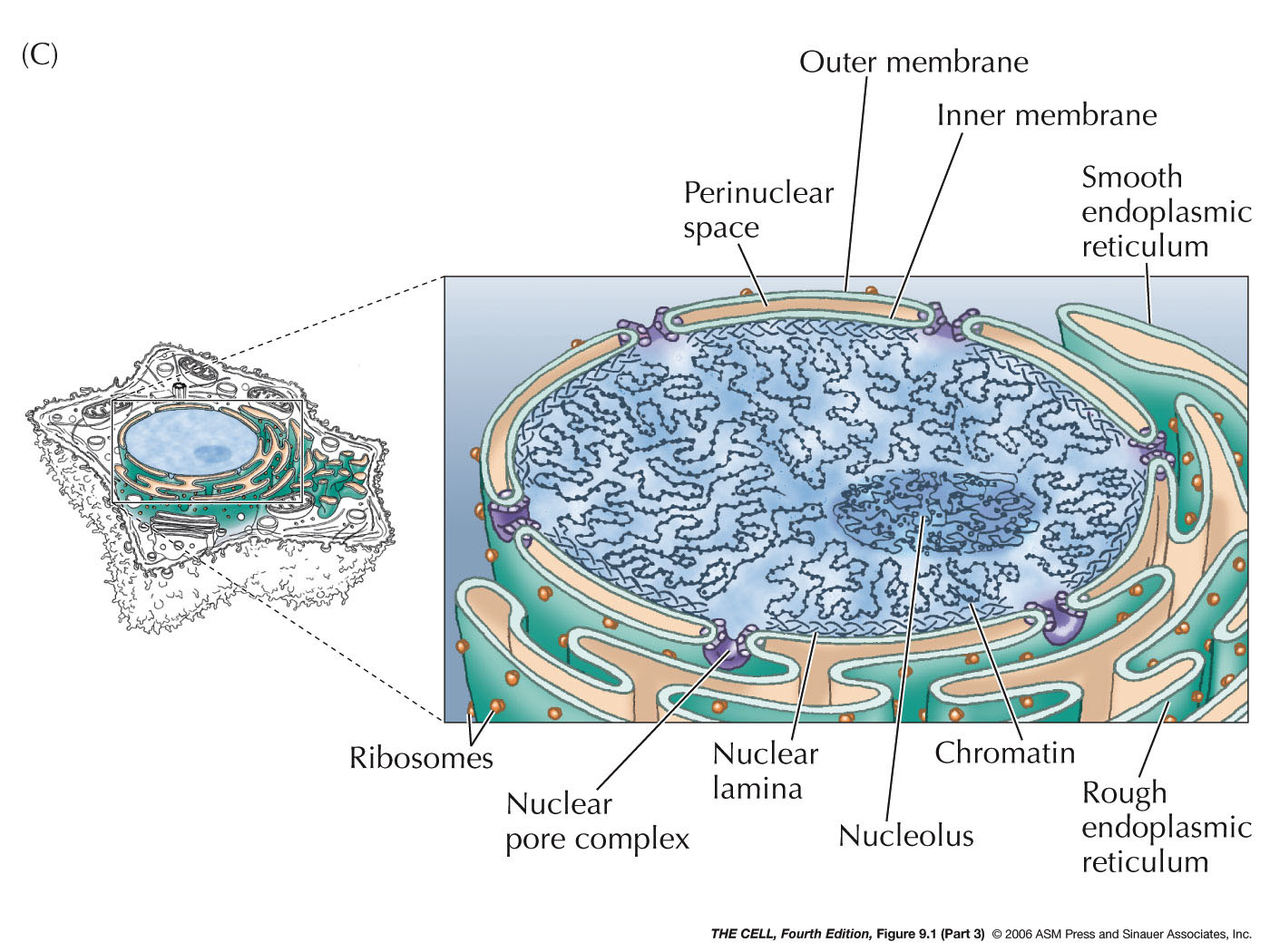
- The
Nuclear
Membrane (Nuclear Envelope): This membrane
separating the nucleus from the cytoplasm is actually a
double membrane
with pores and is attached to the nuclear lamella on the
inside. The outer of the two membranes is continuous with the
endoplasmic reticulum and has all the
characteristics of the ER (including attached
ribosomes). The two membranes have the same phospholipid
bilayer structure seen in the plasma membrane.
|
 |
- The Nuclear Lamina: Just inside the
nuclear membranes is a network of fibrous proteins
(lamins) called the nuclear lamina. Lamins are a type of
intermediate filament proteins (we will see more
examples when we take up the cytoskeleton).
These proteins form tertiary and quaternary
structure (coiled coil and higher order
structures). The lamina binds to the inner nuclear
membrane and to chromatin (H2A and H2B histones).
|

 |
- Nuclear Pore Complex:
While diffusion of small, uncharged molecules is
possible across the phospholipid bilayer,
macromolecules, polar molecules, and ions must enter and
leave the nucleus through nuclear pores.
- Nuclear
Pore Structure: A nuclear pore
joins the inner and outer nuclear membranes and each pore includes
numerous (up to 50 in vertebrates) proteins called
nucleoporins. With the electron microscope a pattern
of eight protein
structures in radial symmetry around the central
channel can be seen. The pore also has a central
ring and a ring on both the cytosolic and nuclear
sides of the pore. Protein filaments extend outward
from the two surface rings.
|
|
|
|
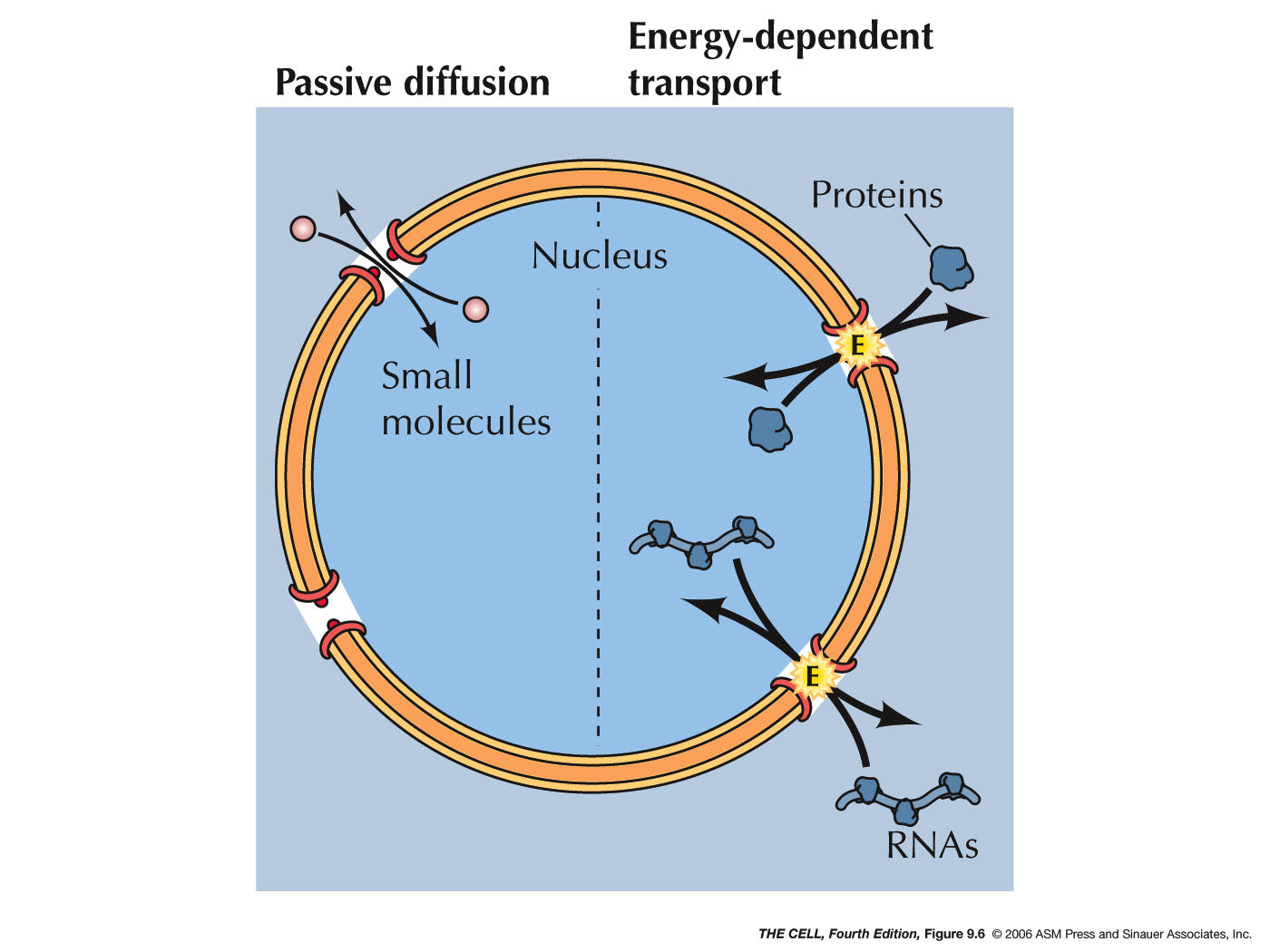 |
- Passive
Diffusion: Smaller proteins (20-40 kd) and
small molecule can freely pass through the open
pores in either direction by simple diffusion.
- Active
Transport: Macromolecules (like larger
proteins and RNAs) and ribonucleoprotein particles
(like preribosomal subunits) must be actively
transported into and/or out of the nucleus through
the pores. Proteins that must enter the nucleus
(they are all made in the cytoplasm) include all of
those involved in DNA and RNA metabolism we studied
earlier (histones, DNA and RNA polymerases, other
replication enzymes, transcription factors, RNA
processing enzymes, plus others). While many
proteins enter the nucleus, some are shuttled back
and forth so must also be able to leave the nucleus.
RNAs made by transcription must be able
to leave the nucleus.
|
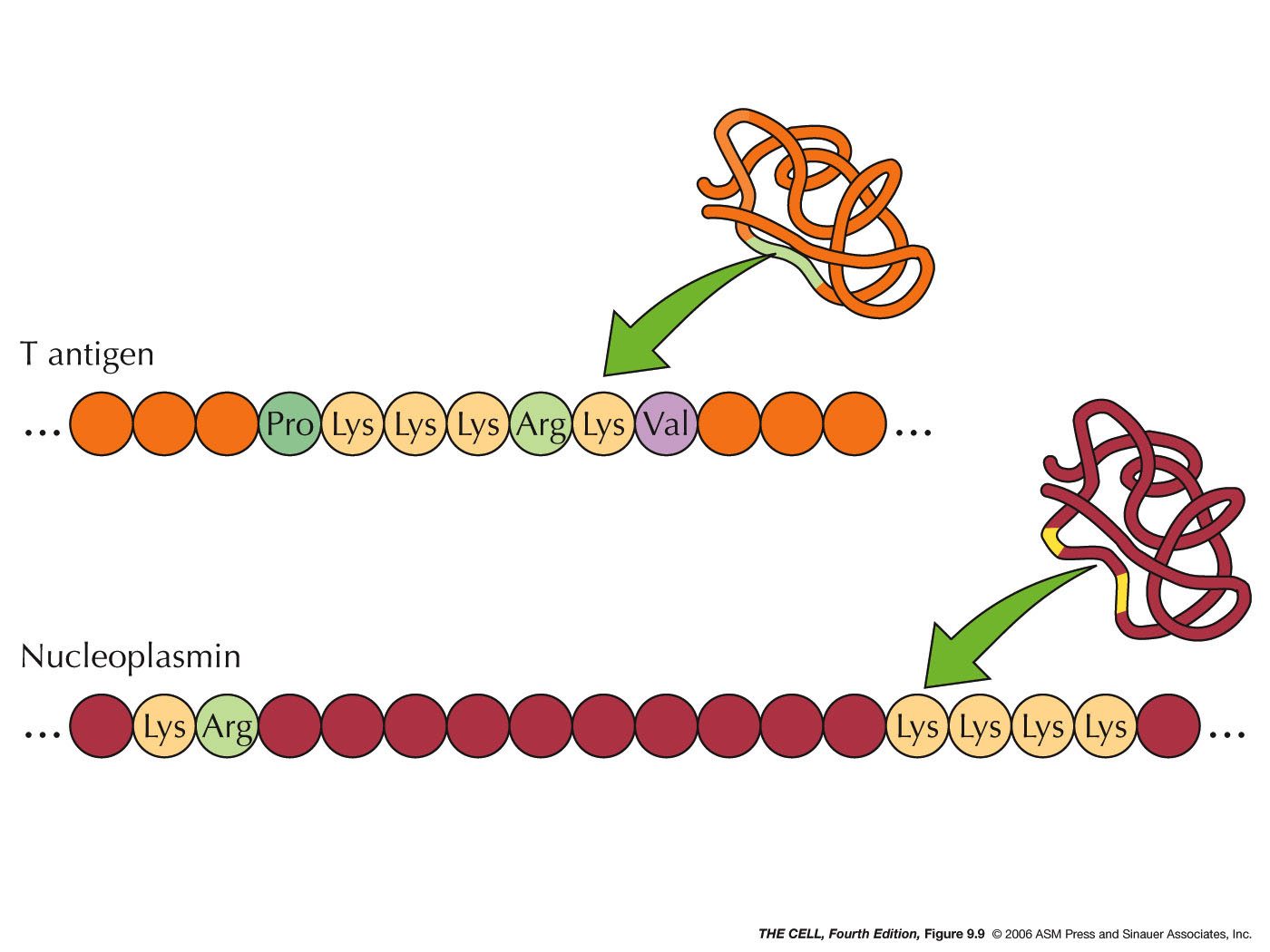
- Protein Entry/Exit: A protein that
is destined to enter the nucleus is marked with a
short amino acid sequence called a nuclear localization
signal (NLS) which is recognized by a
nuclear transport receptor (protein). These short
nuclear localization signal sequences may be
consecutive amino acids or bipartite.
|
 |
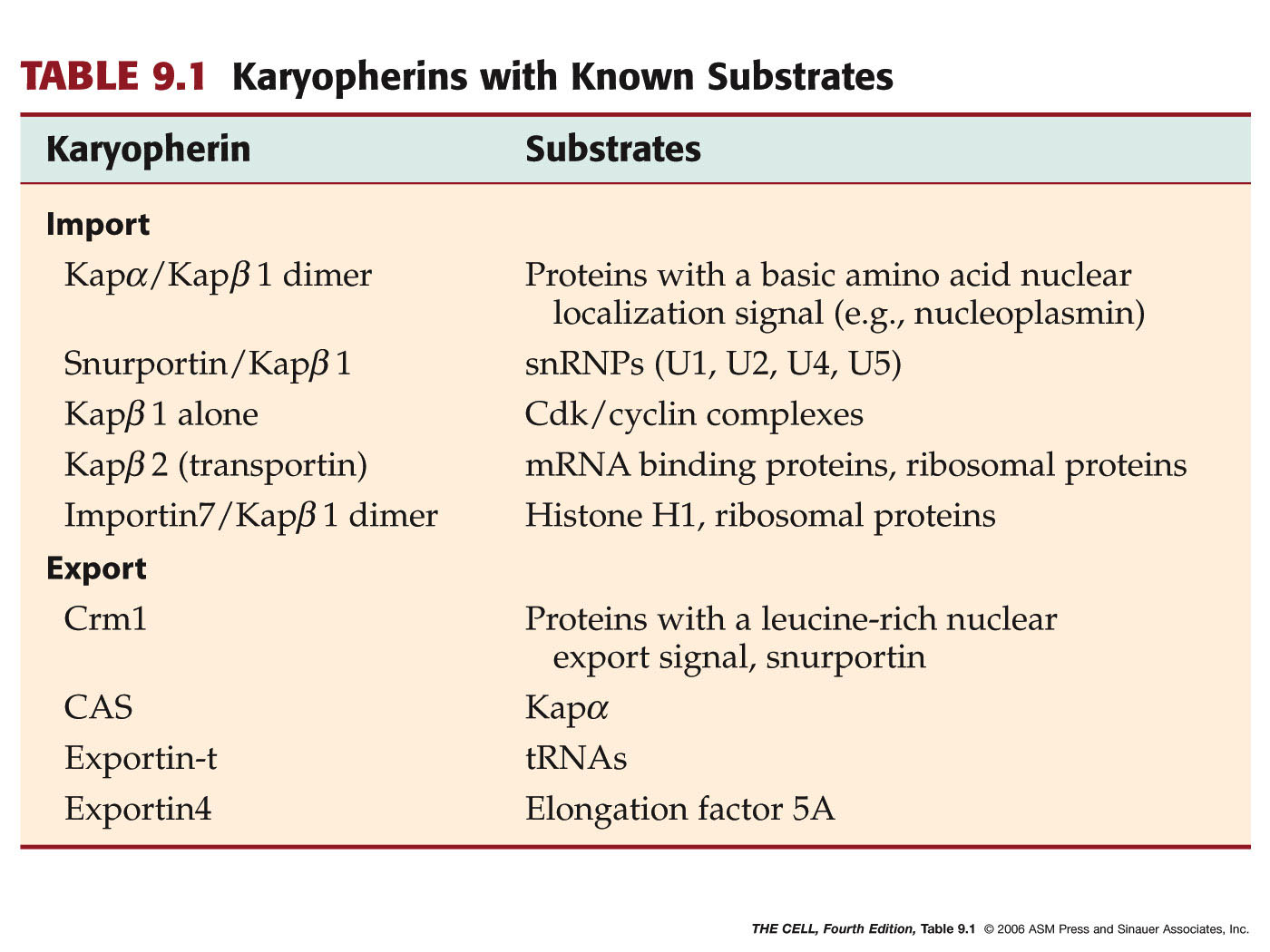
- Karyopherins--an
Example of a Nuclear Transport Receptor:
Karyopherins come in two varieties: importins
and exportins (guess which one imports
macromolecules into the cell and which one
exports). Importins and exportins bind to a
protein to be imported or exported and cross the
nuclear pore with that protein.
|
 |
  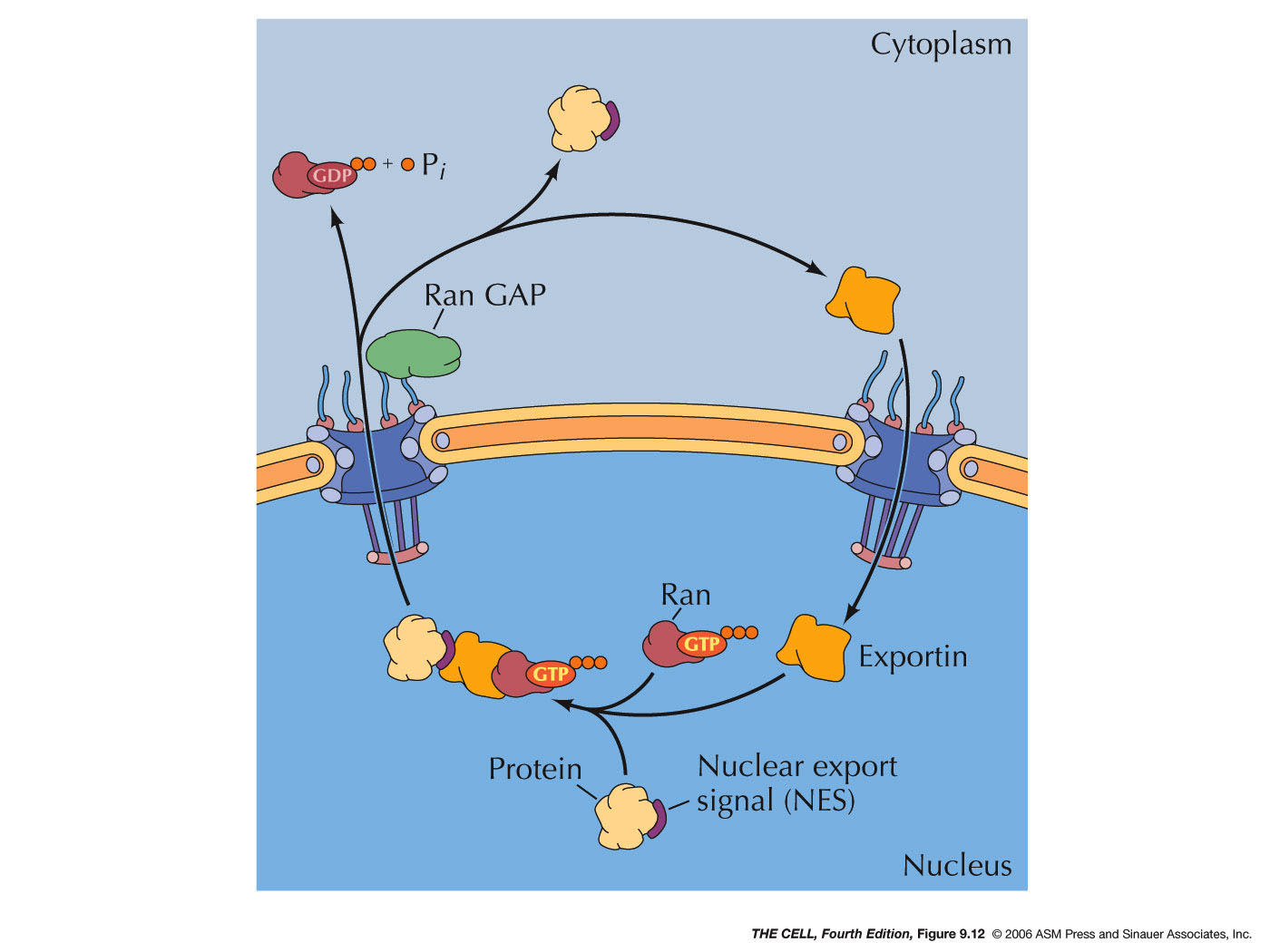 |
- Regulation
of Entry/Exit by the Ran Protein:
- Ran
proteins: Ran proteins
(RAs-related nuclear proteins) are GTP binding
proteins that regulate the entry/exit of other
proteins into/out of the nucleus. The GTP can
be hydrolyzed to GDP and P by an enzyme on the
cytosolic side of the nuclear membrane.
|
- Ran GAP: This
hydrolyzing
enzyme
is called Ran GAP (Ran GTPase-activating
protein).
- Ran GEF: An
enzyme
that
exchanges the GDP bound to Ran for GTP is
localized on the inside of the nuclear
membrane. This enzyme is called Ran GEF (Ran
guanine nucleotide exchange factor).
Therefore, Ran-GTP is in high concentration
inside the nucleus and Ran-GDP is in high
concentration outside the nucleus.
|
- Importins and Exportins: These
are
also
important in this transport process.
- Protein
Import: Import of a protein
begins with the binding of an importin to
the nuclear localization sequence of the
"cargo" protein (the protein to be
imported). This complex binds to nuclear
filaments on the outside of the nuclear pore
and the importin-cargo complex is
transported through the pore. Inside the
nucleus, Ran-GTP binds to the importin and
this binding displaces the cargo, releasing
it inside the nucleus. The Ran-GTP-importin
complex is then transported back through a
pore to the outside where Ran GAP hydrolyzes
the GTP to GDP releasing the importin to be
reused. (The Ran-GDP itself must be
transported by its own transport mechanisms
back into the nucleus where it will be
quickly converted into Ran-GTP by Ran GEF,
replacing its GDP with a GTP.)
|
- Protein
Export: Export occurs by a
similar mechanism except that inside the
nucleus Ran-GTP binding to exportin promotes
the binding of the cargo protein. When this
Ran-GTP-exportin-cargo complex is
transported out of the nucleus, Ran GAP
converts the GTP to GDP. This induces the
release of the cargo protein and the
dissociation of the exportin. (Note: Both
entry and exit require GTP, that is, they
require energy, that is, they are examples
of active transport.)
|
- RNA Entry/Exit: Like the transport
of large proteins across the nuclear membrane, the
import and export of RNAs in and out of the
nucleus is an active (energy-requiring) process.
Ran-GTP dependent importins and exportins
transport most tRNAs, rRNAs, and snRNAs
(similar to protein transport). mRNAs however are
transported by other proteins and apparently do
not require Ran. snRNAs that are part of snRNPs
(used in splicing) are actually transported out of
the nucleus, associate with protein, then the
completed snRNPs are transported back into the
nucleus as a result of nuclear localization
signals on the protein. (We will look at rRNA transport
below under the nucleolus.)
|
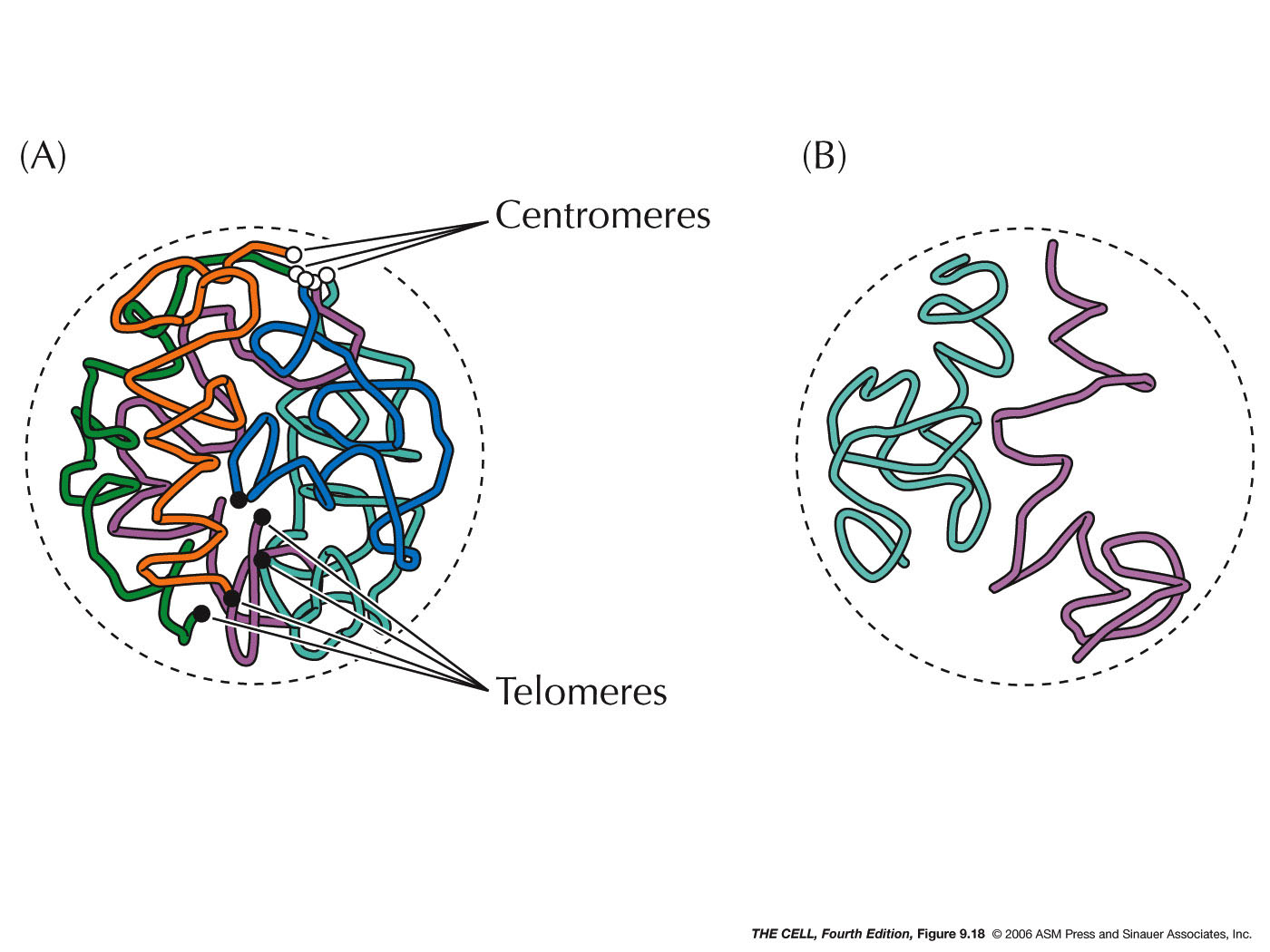
- Organization within the
Nucleus: The nucleus, contrary to earlier
thinking, is not a homogeneous material nor are the
chromosomes randomly distributed in the nucleus. It
shows organization and compartmentalization.
- Chromatin:
This is the stuff chromosomes are made of.
- Chromatin
Condensation during Interphase:
|
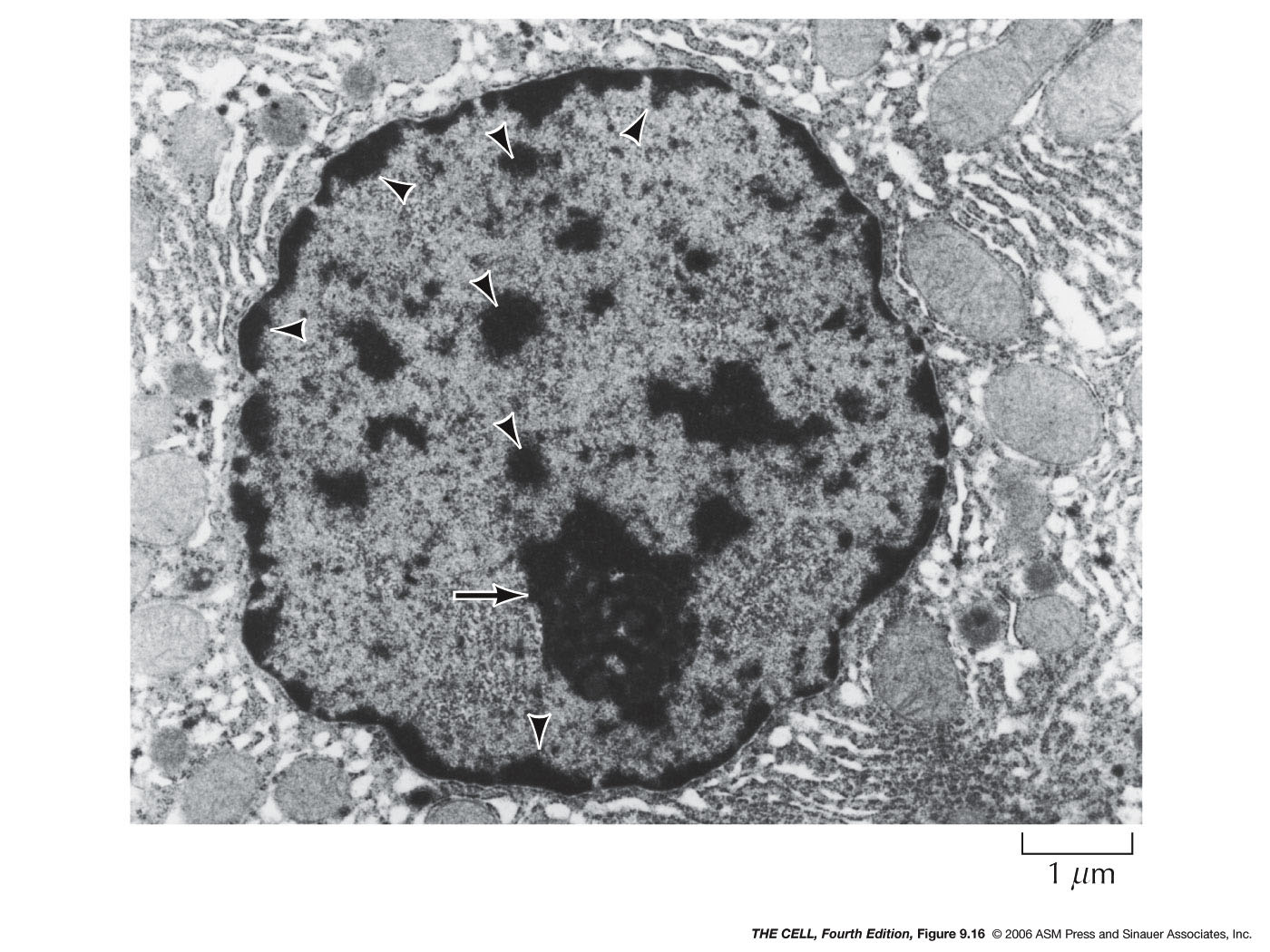
- Heterochromatin:
Chromatin that is condensed during interphase is
called heterochromatin. Heterochromatin is of two
varieties:
|
 |
- Constitutive
heterochromatin is never transcribed
and is condensed in all cells (i.e., satellite
DNA and centromere DNA).
- Facultative
heterochromatin is condensed in
some tissues or at some times but may be
decondensed in other tissues or at other times
(transcriptionally active DNA is decondensed).
|
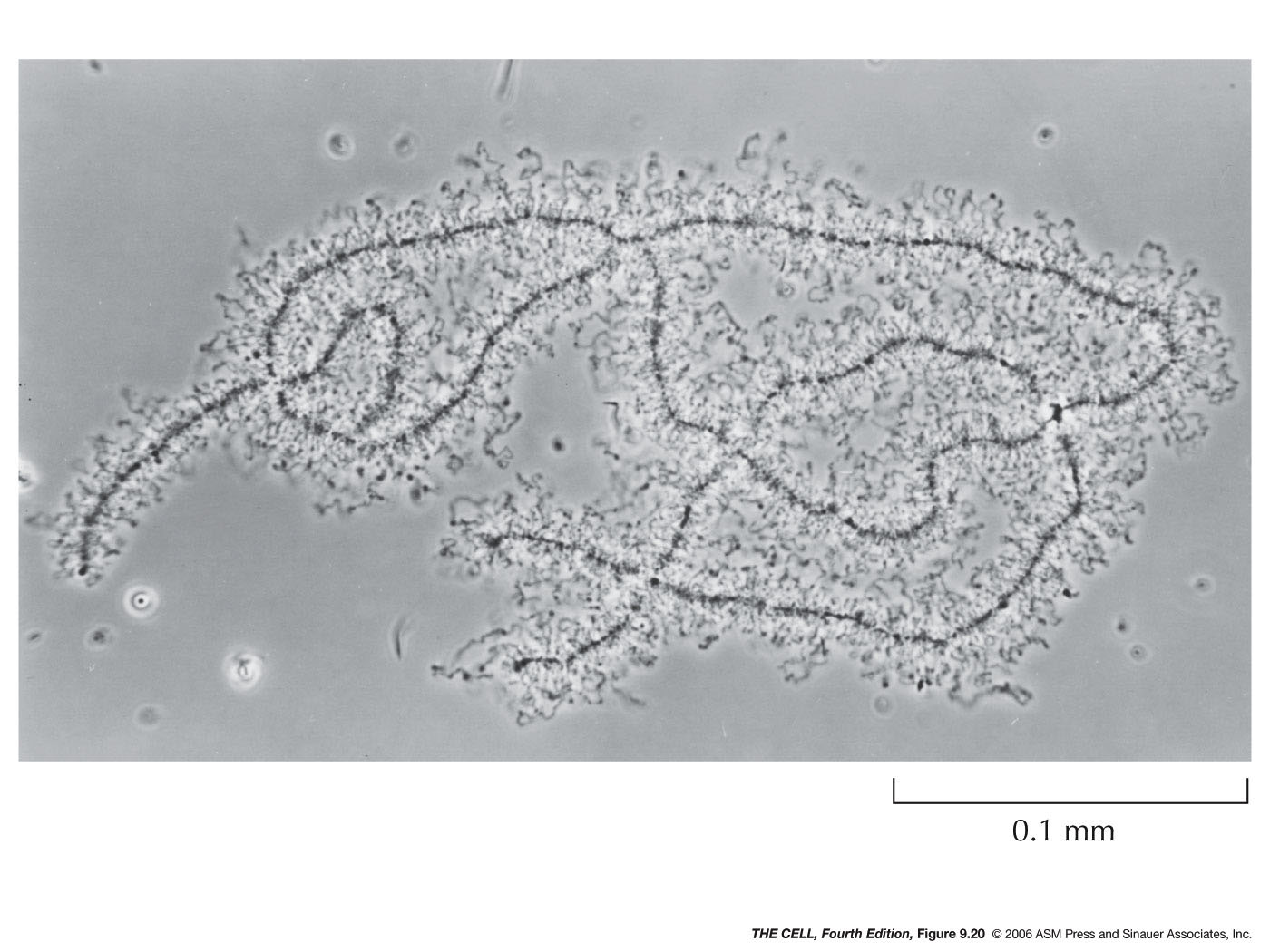
- Euchromatin:
Chromatin that is not condensed during interphase
is called euchromatin.
|
 |
- Localization within
the Nucleus: Certain features and
processes are not randomly distributed in the nucleus
but localized to an area.
|
|
|
 |
|
|
 |
|
|
 |
- The
Nucleolus,
rRNA Processing and Transport, Ribosome Assembly and
Transport: The nucleolus is the site of 45S
pre-rRNA transcription and processing and the site where
ribosomes are assembled. The nucleolus is not membrane
enclosed.
|
- The
NOR: The nucleolus is a structure which is
formed by the aggregated tandemly repeated
genes for the 45S pre-rRNA. In humans, these
genes are on chromosomes number 13, 14, 15, 21, and 22
and make up the nucleolar organizer region (NOR).
|

|
|
|
|
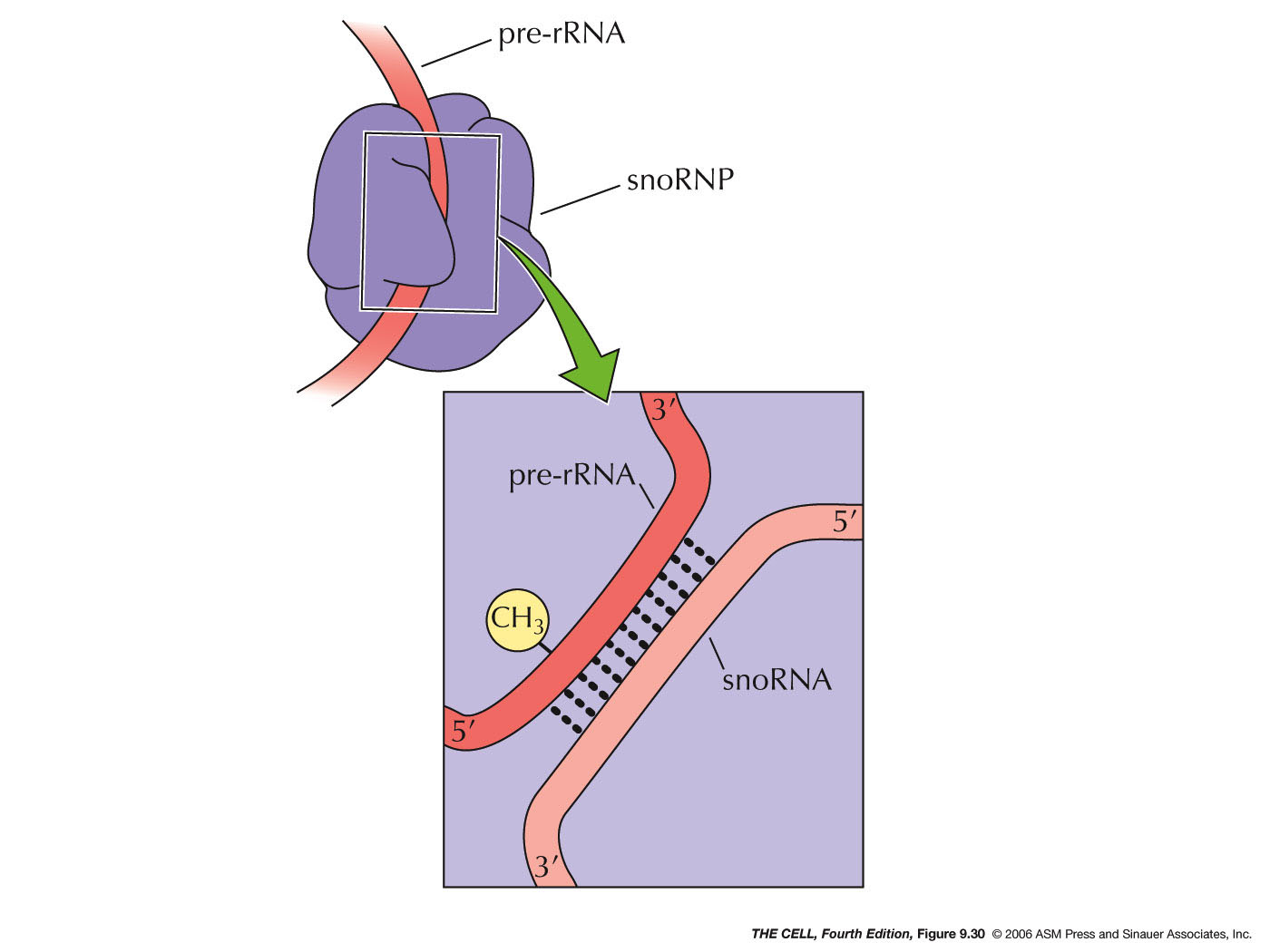
- rRNA Chemical Modification: Also in
the nucleolus rRNA is chemically modified,
especially by methylation.
|
|
|
 |
- 5S rRNA: 5S
rRNAs
are
transcribed outside of the nucleolus by RNA
polymerase III but are assembled into the ribosomes
in the nucleolus.
- rProteins:
The proteins of the ribosome are made like all other
proteins (transcribed by RNA polymerase II and the
transcript is translated in the cytoplasm). They
must then enter the nucleus (see above).
- Ribosome
Assembly and Transport: Ribosomal proteins
begin to aggregate with the 45S pre-rRNA before its
transcription is finished. More than half of them
are bound before this rRNA is completely cleaved.
When these proteins have bound, the 40S preribosomal
subunit (with the 18S rRNA) and 60S preribosomal
subunit (with the 28S, 5.8S, and 5S rRNAs) are
exported from the nucleus through nuclear pores
where they become mature 40S and 60S ribosomal
subunits ready to take part in translation.
|
 Home
Home

















